Research
Mass spectrometry-based methods to study protein amyloid formation
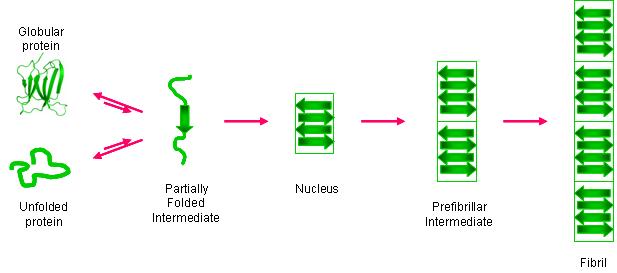
An increasing number of proteins are known to form amyloid fibrils in vivo, and the formation of these fibrils is implicated in several diseases (e.g. Alzheimer’s, Parkinson’s). One of these proteins, β-2-microglobulin (β2m), can form amyloid fibrils, and these fibrils are the main pathogenic process underlying dialysis-related amyloidosis (DRA). Like other amyloid systems, β2m fibril formation proceeds by partial protein unfolding, subsequent oligomerization, and eventual elongation to form mature fibrils. While aspects of general amyloid formation are understood, molecular-level information about the early stages of the amyloid reaction is lacking for almost all amyloid systems; however, this information is critical for the rational design of therapeutics against amyloid diseases. We strive to obtain amino acid-level information of the unfolding and oligomerization of β2m prior to fibril formation by developing mass spectrometry-based methods with the necessary temporal and spatial resolution. We are particularly interested in the role that Cu(II) plays in the amyloid formation of β2m. In general, the methods that we develop not only provide insight into β2m’s amyloidosis but are also general enough to work for other amyloid systems.
Mass Spectrometric Imaging of Nanomaterial Distributions and their Biological Effects
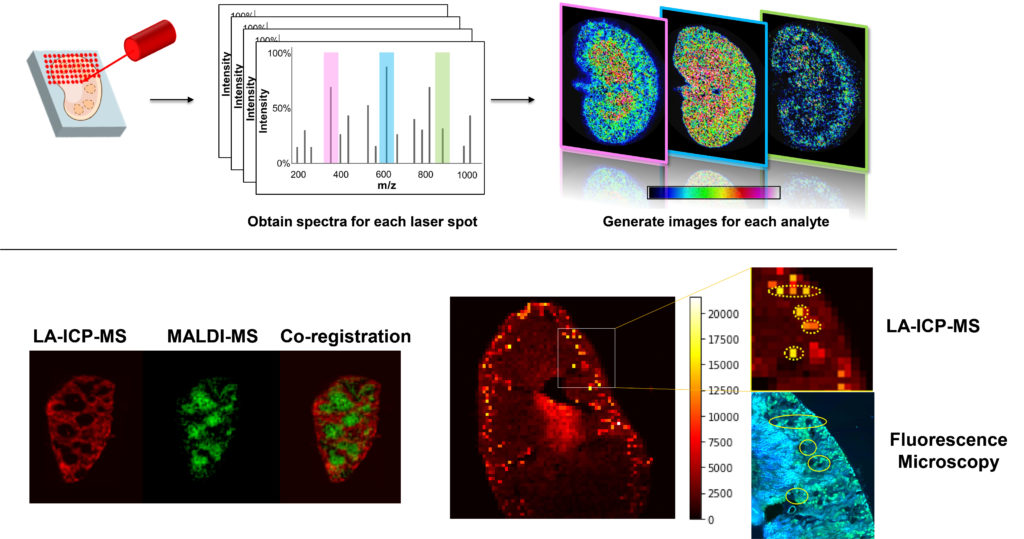
We are also developing new mass spectrometry-based approaches to track, measure, and image nanomaterials in complex samples. Nanomaterials are increasingly being investigated as therapeutic delivery systems, and effective development of such delivery agents requires the ability to monitor them in cells, tissues and organs. Moreover, while these delivery systems may effectively deliver their therapeutic cargo, it is also important to understand any biochemical effects the delivery systems themselves might have. We are developing new multimodal imaging tools that combine MALDI-MS imaging, laser ablation ICP-MS imaging, and microscopy to better understand the fate, transport, bioavailability, and biochemical effects of such nanomaterial delivery systems.
Higher-order Structure of Protein Therapeutics and Epitope Mapping
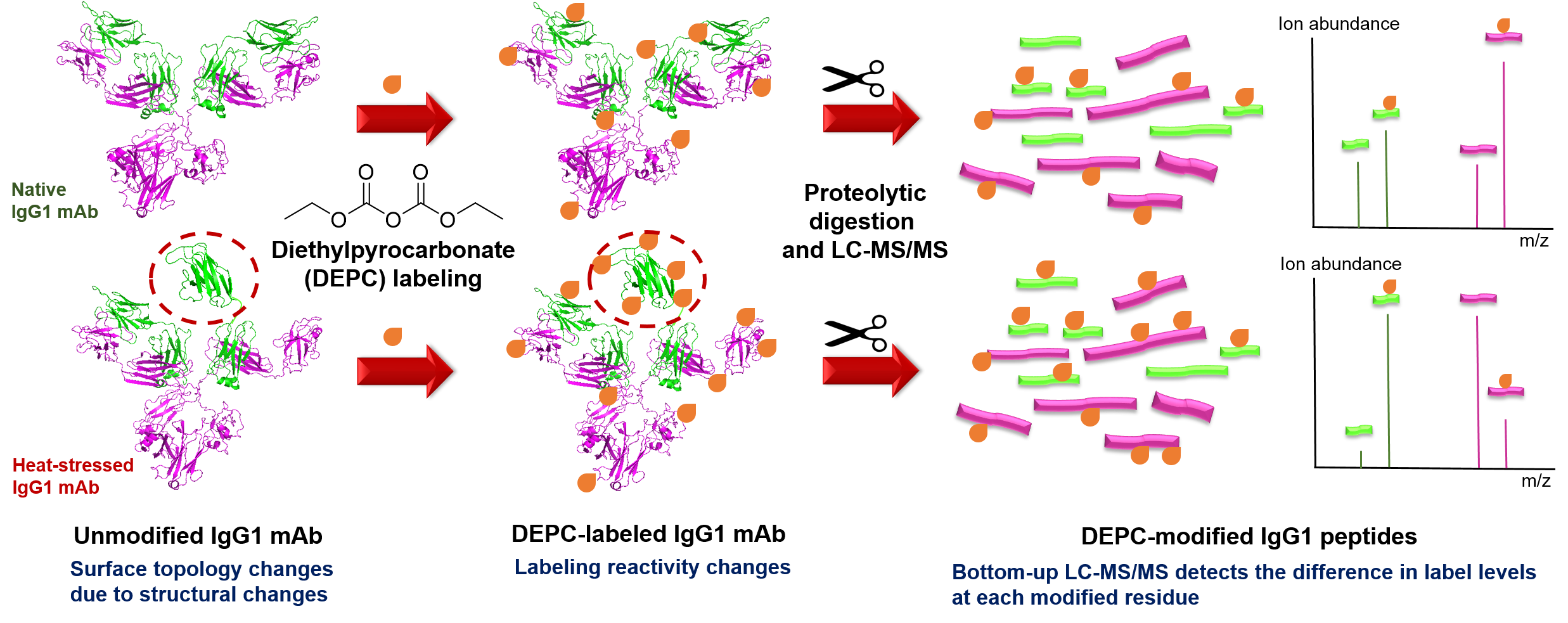
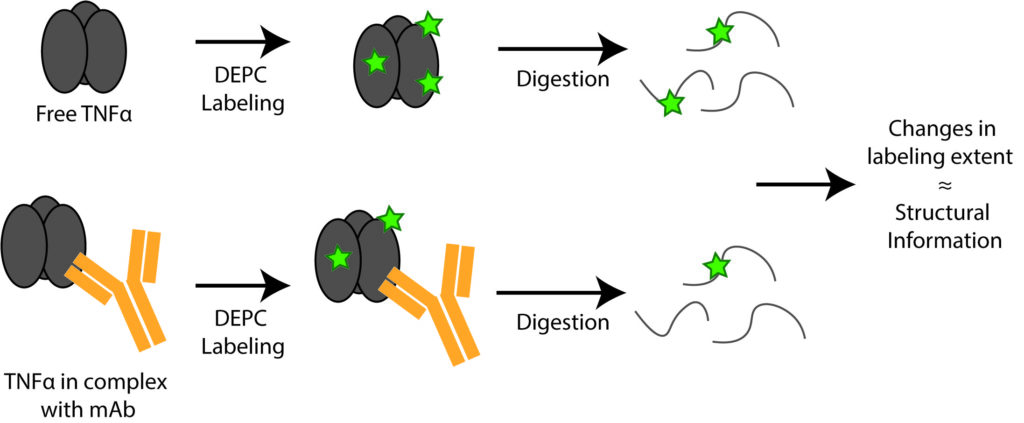 Protein therapeutics account for about one-third of the late-stage drug development pipeline. The higher order structure (HOS) and binding interactions of these therapeutics are essential to monitor to ensure their efficacy and safety. HOS changes can lead to reduced stability, loss of efficacy, and possible immunogenicity. We have explored the use of covalent labeling and mass spectrometry (CL-MS) as a means to detect HOS changes to therapeutic proteins and to identify the epitopes they recognize on their corresponding antigen. Diethylpyrocarbonate (DEPC) CL-MS provides moderate structural resolution in a relatively efficient manner.
Protein therapeutics account for about one-third of the late-stage drug development pipeline. The higher order structure (HOS) and binding interactions of these therapeutics are essential to monitor to ensure their efficacy and safety. HOS changes can lead to reduced stability, loss of efficacy, and possible immunogenicity. We have explored the use of covalent labeling and mass spectrometry (CL-MS) as a means to detect HOS changes to therapeutic proteins and to identify the epitopes they recognize on their corresponding antigen. Diethylpyrocarbonate (DEPC) CL-MS provides moderate structural resolution in a relatively efficient manner.